Normal mode analysis of macromolecular systems with the mobile block Hessian method
Abstract
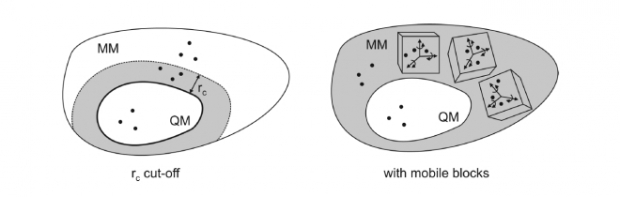
Until recently, normal mode analysis (NMA) was limited to small proteins, not only because the required energy minimization is a computationally exhausting task, but also because NMA requires the expensive diagonalization of a 3N(a) x 3N(a) matrix with N-a the number of atoms. A series of simplified models has been proposed, in particular the Rotation-Translation Blocks (RTB) method by Tama et al. for the simulation of proteins. It makes use of the concept that a peptide chain or protein can be seen as a subsequent set of rigid components, i.e. the peptide units. A peptide chain is thus divided into rigid blocks with six degrees of freedom each.
Recently we developed the Mobile Block Hessian (MBH) method, which in a sense has similar features as the RTB method. The main difference is that MBH was developed to deal with partially optimized systems. The position/orientation of each block is optimized while the internal geometry is kept fixed at a plausible - but not necessarily optimized - geometry. This reduces the computational cost of the energy minimization. Applying the standard NMA on a partially optimized structure however results in spurious imaginary frequencies and unwanted coordinate dependence. The MBH avoids these unphysical effects by taking into account energy gradient corrections. Moreover the number of variables is reduced, which facilitates the diagonalization of the Hessian.
In the original implementation of MBH, atoms could only be part of one rigid block. The MBH is now extended to the case where atoms can be part of two or more blocks. Two basic linkages can be realized: (1) blocks connected by one link atom, or (2) by two link atoms, where the latter is referred to as the hinge type connection. In this work we present the MBH concept and illustrate its performance with the crambin protein as an example.