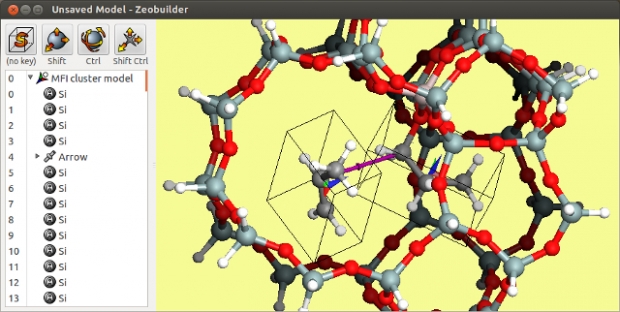
We generously share our software under the conditions of the GNU GPLv3 license, but we need your help to guarantee the continuity of these projects. Therefore we kindly ask a few favors in return for the use of our software:
- Register for free.
- Cite the appropriate papers when you use our software for your research or to prepare publications.
- Provide feedback by using the mailing list of the software package.
Read our flyer (version 07/2012)