Unraveling the vibrational fingerprint of guest-loaded nanoporous materials through computational derivation and partitioning of IR and Raman spectra
Unraveling the vibrational fingerprint of guest-loaded nanoporous materials through computational derivation and partitioning of IR and Raman spectra
Promotor(en): V. Van Speybroeck /19SPEC05 / SpectroscopyNanoporous materials, which are crystalline and have pores at the nanometer scale, have a wide range of applications ranging from gas separation and sensing to chemical catalysis. An example is given by zeolites, which consist of aluminosilicate units forming a three-dimensional crystalline network, and represent fascinating materials that are vital in various industries. For example, zeolites play an important role in the storage of thermal energy through the adsorption of water as well as in upcoming technologies to convert biomass or even CO2 into hydrocarbons, which are the building blocks for polymers. Given their economic impact, there is a powerful incentive for the characterization and smart design of new functionalized materials to obtain the best material for a given application.
From an experimental point of view, it is a challenge to obtain insight into the microscopic structure of the material and even more so for the adsorbed guest species. Nevertheless, a wide range of spectroscopic techniques exist to characterize the materials. Recent advances made operando spectroscopic measurements possible, leading to a wealth of information while the system is at work. Vibrational spectroscopic tools, such as infrared (IR) and Raman spectroscopy, are especially well-suited to follow molecules in the zeolite pore system at operating conditions. These techniques allow to generate a fingerprint for the materials and its guest species. Unfortunately, it is extremely difficult to interpret the complex experimental spectra. It is in this respect that computational modeling offers an attractive complementary technique.
Nowadays, molecular simulations are able to accurately reproduce experimental spectra and also allow for the assignation of various spectral features to specific molecular motions. Yet, this assignation is still a challenge in complex systems, such as zeolites containing guest species, even with the help of theoretical techniques. On the one hand the vibrational spectrum needs to be subdivided into vibrational modes to determine the nature of the molecular motions. This becomes increasingly difficult with increasing system size. On the other hand the IR and Raman intensities need to be calculated for both the zeolite framework and the guest species requiring expensive ab initio calculations based on quantum mechanics to accurately determine the electronic structure.
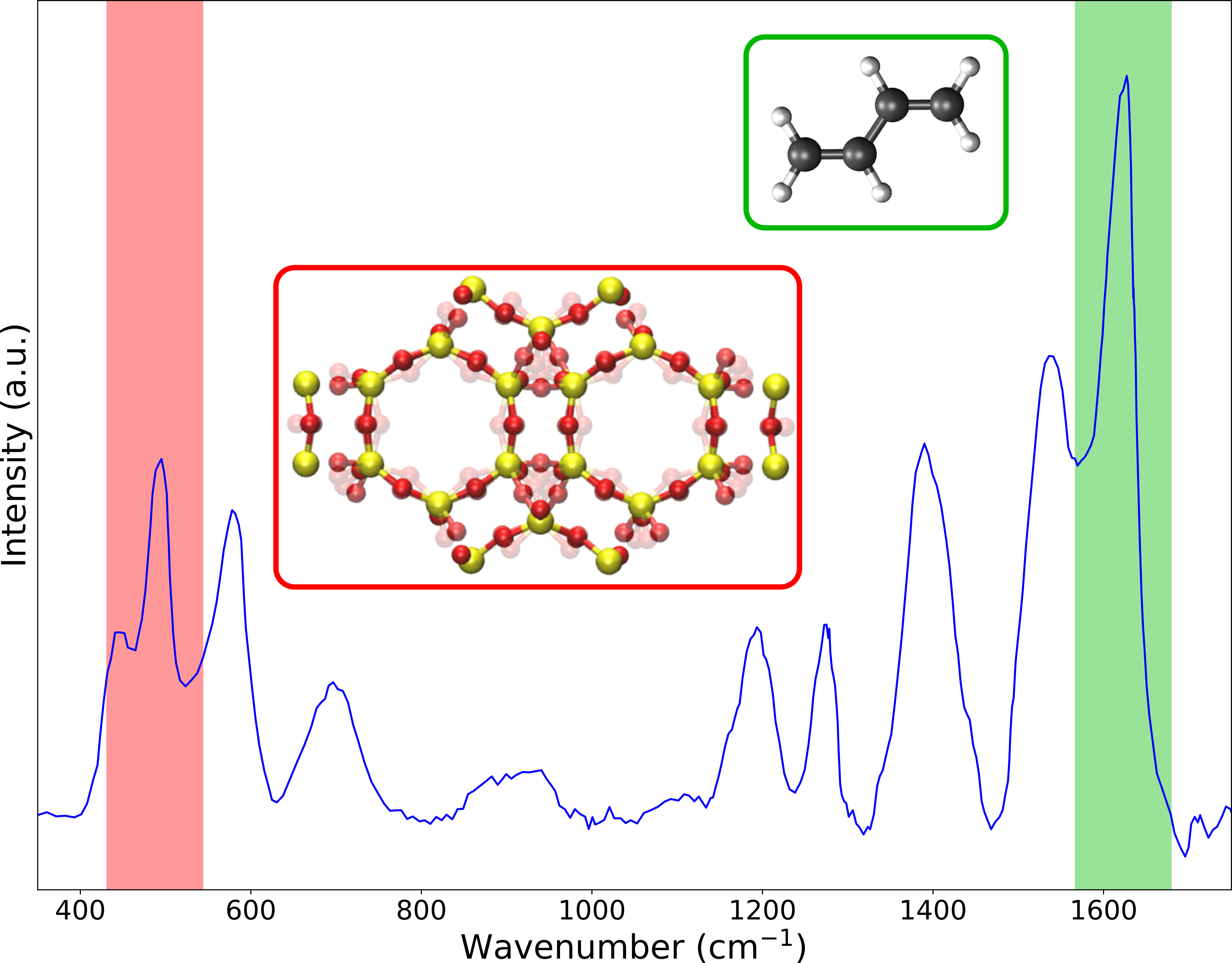
Goal
In this project the vibrational spectrum of zeolite frameworks hosting guest molecules will be constructed and analyzed in detail by means of computational modeling. For that purpose, various tools need to be developed which are able to extract the vibrational modes of the system and determine the IR and Raman intensities of the different components. Both tasks rely on an accurate prediction of the potential energy surface (PES) which will initially be constructed based on methods whereby the Schrödinger equation needs to be solved. However, as these methods are computationally expensive, force fields might be used to speed up the calculations.
The first objective is to determine the vibrational modes of the host-guest system. There exist two fundamentally different approaches to perform this task: either based on static or dynamic simulations. The static approach is used more often due to its lower computational cost. It starts from the equilibrium structure, which is a minimum on the PES, and approximates the surrounding PES by means of a parabola, the so-called harmonic approximation. As a result, Newton’s equations of motion can be solved in terms of uncoupled harmonic oscillators at fixed frequencies. The corresponding vibrational modes are denoted as normal modes and this technique is labeled as normal mode analysis (NMA). However, in nanoporous materials hosting guest molecules, the equilibrium configuration is hard to find and does not account for the mobility of the guest molecules. Therefore, molecular dynamics (MD) simulations are required to sample the PES, in which Newton’s equations of motion are integrated yielding a trajectory of the atomic positions as a function of time. This trajectory implicitly contains the vibrational modes of the system, which need to be extracted by numerical techniques in order to identify the origin of the experimentally observed vibrational fingerprints. In literature, there already exist various procedures to obtain the modes from an MD trajectory, such as Principal Component Analysis (PCA). In this project, these techniques will be to implemented in Yaff, an in-house software package for performing and post-processing molecular dynamics simulations. Furthermore, the student will also be encouraged to propose extensions to NMA and PCA that promote more localized vibrational modes, allowing more straightforward interpretation of the corresponding molecular motions.
A second objective then focusses on computing the IR and Raman activity corresponding with each vibrational modes. In case of normal mode analysis, the normal modes correspond to harmonic vibrations at fixed frequency. As a result, each normal mode corresponds with a single peak in the IR/Raman spectrum for which the corresponding intensity can be computed as the amplitude of the fluctuating dipole moment and polarizability. In the case of PCA modes, one cannot necessarily assign each mode to a single frequency peak. However, one can still compute to corresponding intensity for each mode. Together with the spectral decomposition of each vibrational mode (i.e. what frequencies are present in each mode), this information will be used for the interpretation of the experimental spectra and the corresponding assignation of modes.
As a final objective, the IR and Raman spectra of host-guest systems will also be computed directly as the Fourier transform (i.e. spectral decomposition) of the time-dependent dipole moment and polarizability respectively. The calculation of these quantities of the complete unit cell by means of ab initio methods has become possible since the development of the modern theory of polarization. These ab initio spectra can serve as a theoretical counterpart of the experimental spectra to assess the quality of the various computational models used throughout the thesis. Unfortunately, this does not allow for a straightforward assignation of each peak in the resulting spectra. Nevertheless, different localization techniques have emerged which could solve the problem, such as the use of Wannier centers. Although these techniques have proven their worth for the interpretation of vibrational spectra of solutions, they have never been used on nanoporous systems containing guest molecules. In this project, the performance of such localization schemes within zeolite systems will be assessed and applied.
Aspects
Master of Science in Engineering Physics: This thesis subject is closely related to the following clusters of elective courses: Nano and Modeling. Engineering aspects: the student will apply advanced molecular simulations and develop new computational tools to understand complex vibrational spectra in zeolites, which is of high industrial importance. Physics aspects: the student will use several techniques (normal mode analysis, ab initio calculations, …) to compute various physical properties (dipole moment, polarizability). Interpreting the results will require a fundamental understanding of quantum mechanics, solid state physics, atomic and molecular physics and statistical physics.
- Study programmeMaster of Science in Engineering Physics [EMPHYS], Master of Science in Physics and Astronomy [CMFYST]ClustersFor Engineering Physics students, this thesis is closely related to the cluster(s) NANO, MODELINGKeywordsVibrational spectroscopy, normal mode analysis, infrared spectrum, raman spectrum, quantum mechanical calculation, Molecular dynamics
